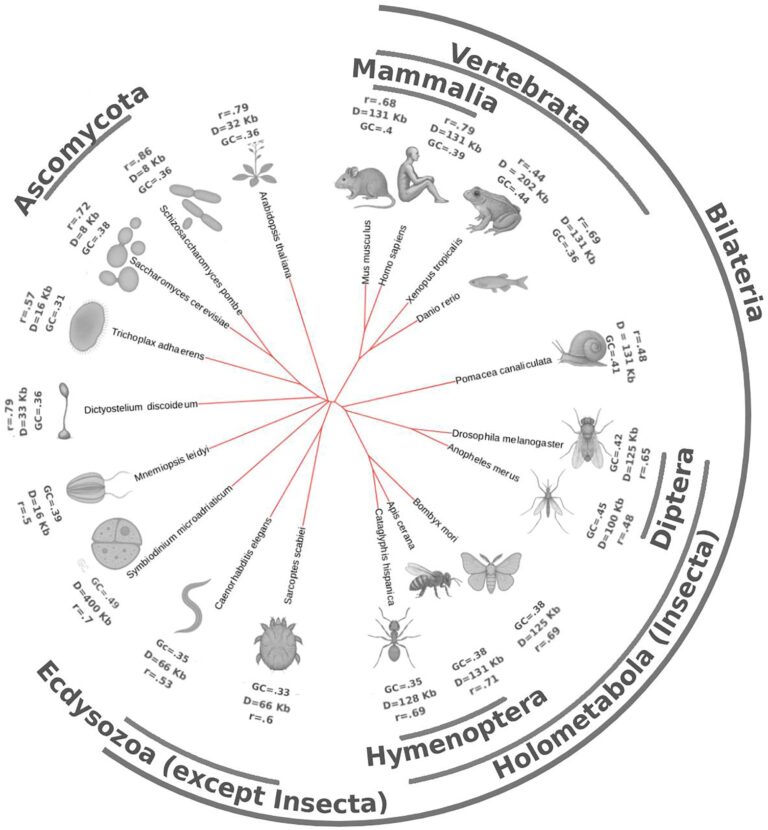
A very curious part of my PhD work on Dictyostelium discoideum chromatin has been published in Nucleic Acid Research this week. My part was to design a 1D extrusion and polymer model that explains why the “bright dots” are happening at the convergent gene positions (CGPs).
After first seeing this fascinating pattern, I hypothesized that cohesin might be dragged to the CGPs by moving polymerase, following up ideas and findings in bacteria by Hugo and Aafke and in mammals by Ed and Aafke. Turns out, simulations only hold if cohesin is significantly slower than polymerase—a strong assumption that requires further confirmation!
Although we found substitutions in ATP binding in the ATPase heads of the Dicty cohesin that might destabilize ATP binding and lead to a slower speed, I believe that further research and evidence will be essential to solving the puzzle.
I look forward to further research on Dicty and cohesin variants to determine whether my hypothesis holds.
Irina V Zhegalova, Sergey V Ulianov, Aleksandra A Galitsyna, Ilya A Pletenev, Olga V Tsoy, Artem V Luzhin, Petr A Vasiluev, Egor S Bulavko, Dmitry N Ivankov, Alexey A Gavrilov, Ekaterina E Khrameeva, Mikhail S Gelfand, Sergey V Razin.
Nucleic Acids Research, Volume 53, Issue 2, 27 January 2025, gkaf006.
DOI: https://doi.org/10.1093/nar/gkaf006